
Structural Variants in Bottlenose Dolphins
- KEYWORDSstructural variants, bottlenose dolphin, population genetics, bioinformatics, Manta, indels, ecotypes
- ADVISORDr. Tom Schultz, Duke University Marine Lab
- WEBSITEhttps://tinyurl.com/y3dw4wex
Overview
This was my summer 2023 project as a Bonaventura Fellow at the Duke Marine Lab. From a high level, studying the underlying genetic mechanisms of speciation and population-level differentiations can provide a window into the past context or future possibility of speciation events. A new avenue for studying variation beyond single-nucleotide polymorphisms (SNPs) comes in the form of stuctural variants (SVs), large chromosomal-level alterations larger than 50 base pairs in size.
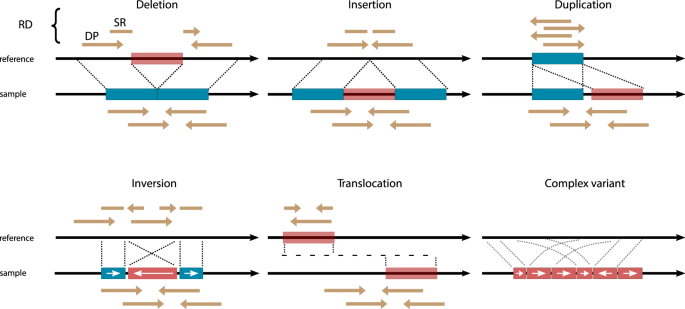
Examples of structural variants. Image from van Belzen et al., 2021.
Our study organism for detection and analysis of structural variants was the bottlenose dolphin. In the western North Atlantic, three ecotypes of common bottlenose dolphins are present - referred to as inshore, offshore, and coastal. The offshore and coastal ecotypes are separated by the continental shelf and have developed distinct phenotypic traits. While previous research using whole genome data of western North Atlantic bottlenose dolphins focused on SNP calling, our study attempts to use compiled short-reads to call SVs and instead provide a foundation for understanding whether SVs play a part in the differentiation and potential speciation between these two distinct ecotypes in addition to the third discrete inshore ecotype.”
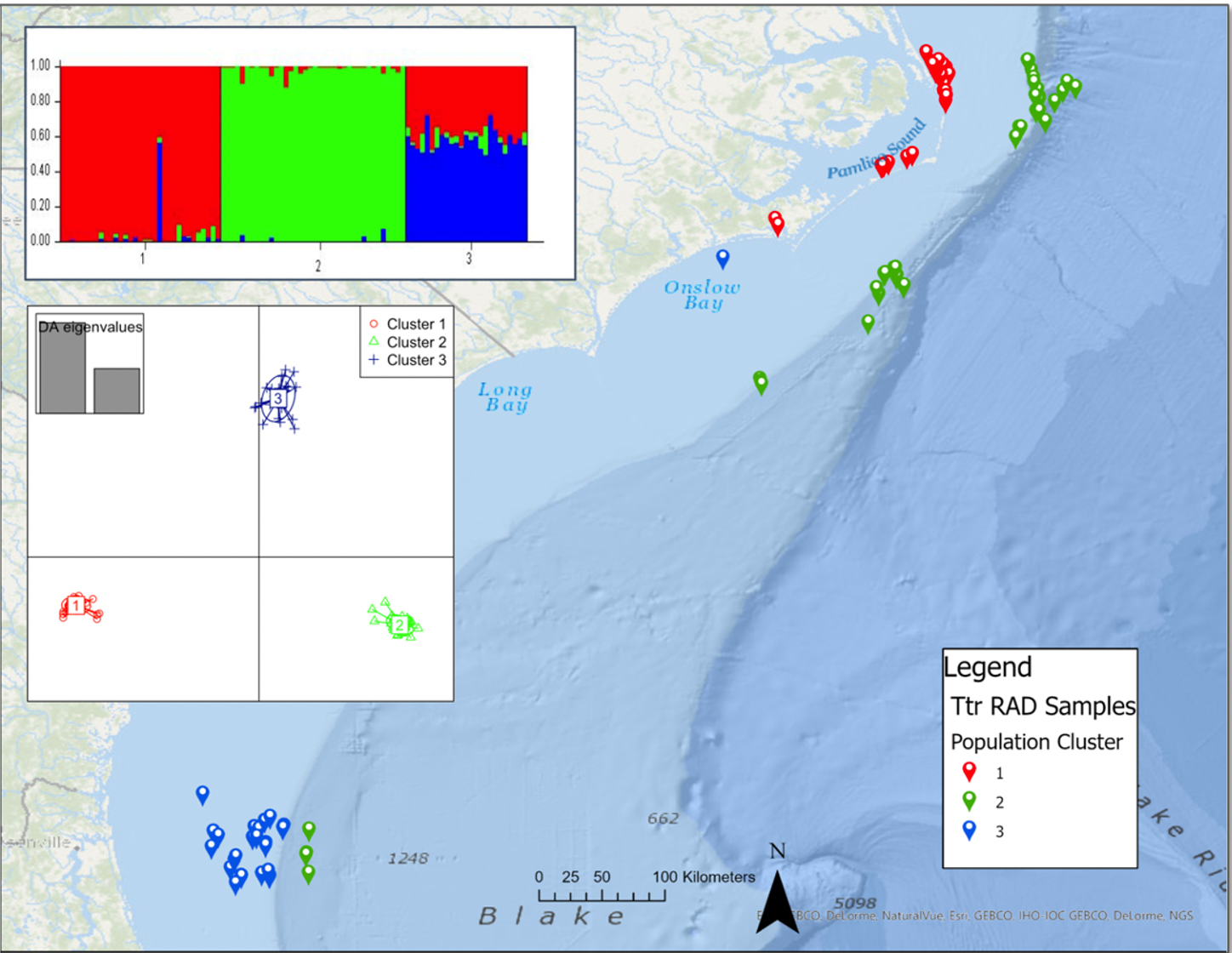
Past work in bottlenose dolphins (see DAPC and STRUCTURE plot here) using SNPs has revealed three genetically distinct ecotypes. Red = inshore, green = offshore, blue = coastal.
Results
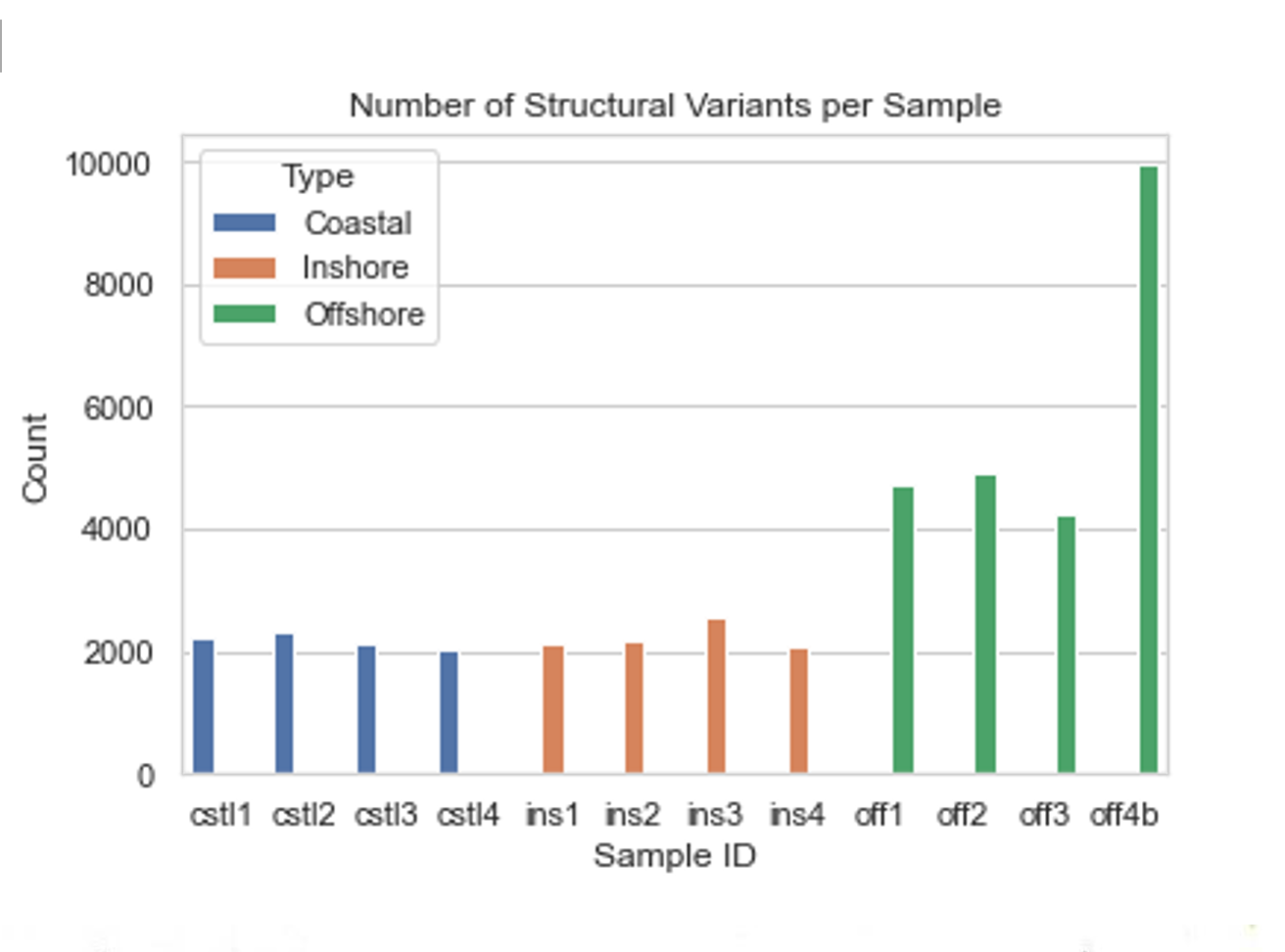
Variant calling was done using Manta software on binary-aligned reads in .bam format. For this project, I had sequences for 12 individuals with 4 from each ecotype present. Prior to filtering, Manta discovered over 32,000 variants (15,520 after filtration for quality/depth). On average, coastal samples contained the least number of variants and insertions and deletions (indels) at 2173 and 900, respectively. Inshore samples averaged slightly more, at 2232 and 920 variants and indels. Offshore samples, however, contained substantially more variants, averaging over twice as many at 5962 variants, of which 2520 were indels.
After initial analysis of called variants, I ran PCA and DAPC to cluster the samples, and just like the a priori ecotype designation which was done with SNPs, we ended up with 4 samples in each of the 3 clusters (inshore, offshore, coastal). This was mostly where I wrapped up analysis over the summer, although I did go hunting through the .vcfs to both get summary statistics about variant sizes and types for a particular individual (not shown here) and to find a few variants of interest for my class lab project. I took Tom (my advisor)‘s course on marine mammal genomics in spring 2024 and did my lab project on genotyping individuals using some 50-100 base pair variants that I called last summer. For more information about this, see my IS paper.
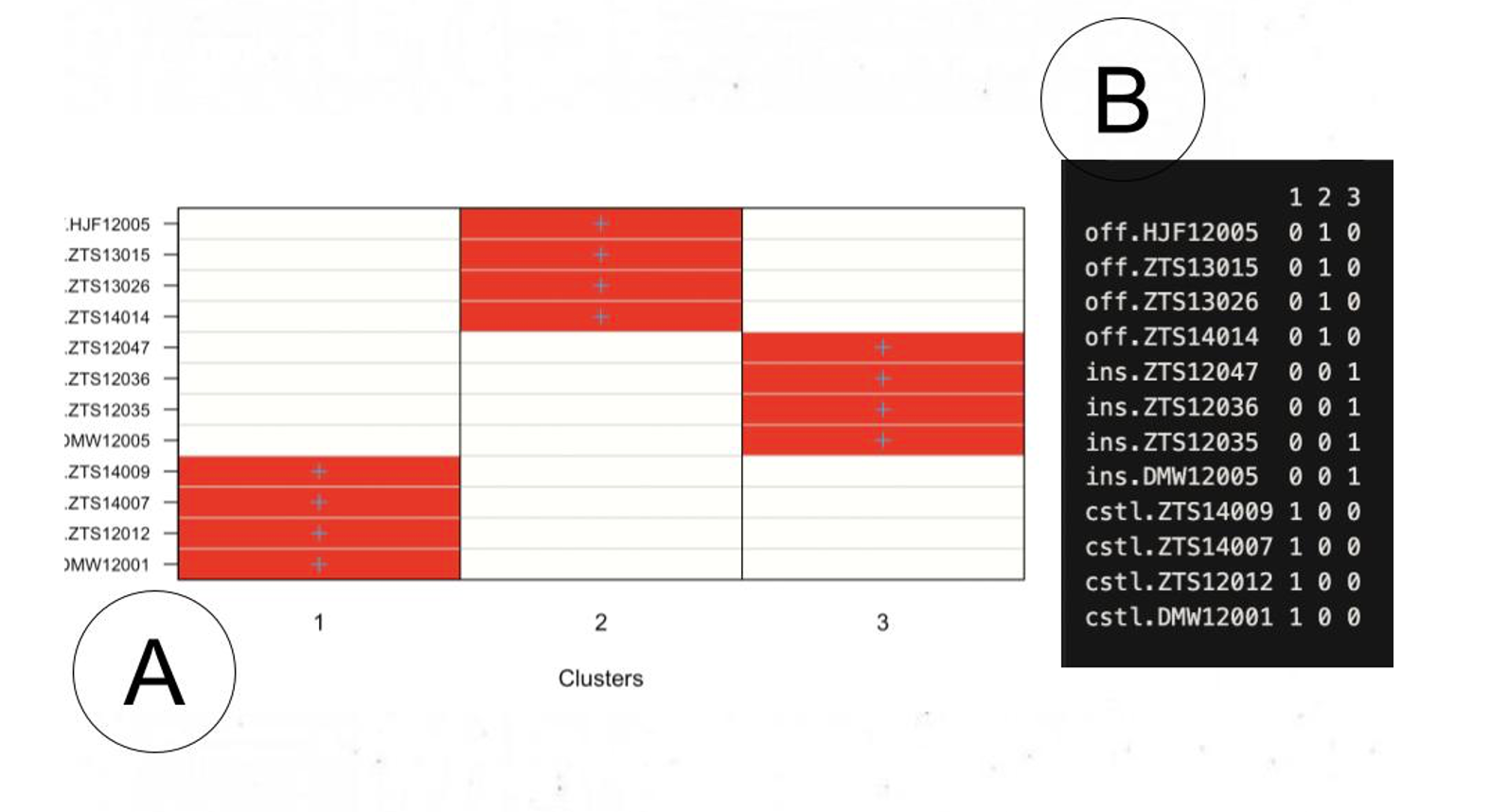
DAPC: Coastal individuals mapped to cluster 1, offshore to cluster 2, and inshore to cluster 3. Sorry, not the best figure.
Future Work
While I’m no longer working on this project, identifying functional polymorphic SVs that could be responsible for ongoing differentiation between dolphin populations is a compelling area of research.